Sustainable Therapeutic Strategies for Controlling Amyloid Aggregation Process Polymer Science
Main Article Content
Abstract
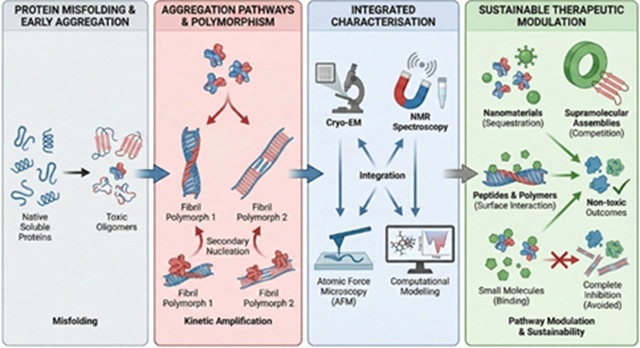
Protein misfolding and aggregation into amyloid assemblies underlie a broad class of neurodegenerative and systemic disorders, including Alzheimer’s, Parkinson’s, and Huntington’s diseases. Although amyloid deposition has long been recognised as a pathological hallmark, increasing evidence indicates that disease progression is driven by pathway-dependent aggregation processes involving transient oligomeric intermediates, fibril polymorphism, and surface-mediated amplification mechanisms, rendering amyloid aggregation both mechanistically complex and therapeutically challenging. Recent advances in experimental biophysics and computational modelling have substantially refined understanding of amyloidogenesis. High-resolution structural techniques, together with kinetic and spectroscopic assays, have clarified how sequence features, environmental conditions, and aggregation history shape the structural and toxic properties of amyloid assemblies. In parallel, atomistic and coarse-grained simulations, multiscale modelling, and data-driven approaches have enabled systematic interrogation of misfolding pathways, energetic landscapes, and kinetic control points that are difficult to access experimentally, while also supporting more efficient experimental design. Against this mechanistic backdrop, therapeutic development has shifted from non-specific aggregate clearance toward precise modulation of aggregation pathways. Emerging strategies emphasise sustainability-oriented principles, including selectivity, reversibility, reduced material complexity, and compatibility with green chemistry. Small molecules, nanomaterials, supramolecular assemblies, peptide-based constructs, and polymeric systems are being developed to bias aggregation trajectories, attenuate secondary nucleation, or destabilise toxic intermediates rather than enforce complete inhibition. This review highlights recent progresses in amyloid aggregation and presents a computationally guided, sustainable framework for disease-specific aggregation control.
Article Details

This work is licensed under a Creative Commons Attribution 4.0 International License.
How to Cite
References
F. Chiti, C. M. Dobson. Protein misfolding, functional amyloid, and human disease. Annu. Rev. Biochem. 2006, 75, 333. https://doi.org/10.1146/annurev.biochem.75.101304.123901 DOI: https://doi.org/10.1146/annurev.biochem.75.101304.123901
T. P. Knowles, M. Vendruscolo, C. M. Dobson. The amyloid state and its association with protein misfolding diseases. Nat. Rev. Mol. Cell Biol., 2014, 15, 384. https://doi.org/10.1038/nrm3810 DOI: https://doi.org/10.1038/nrm3810
P. Ghosh, P. De. Modulation of amyloid protein fibrillation by synthetic polymers: Recent advances in the context of neurodegenerative diseases. ACS Appl. Bio Mater., 2020, 3, 6598. https://doi.org/10.1021/acsabm.0c01021 DOI: https://doi.org/10.1021/acsabm.0c01021
E. Agorogiannis, G. Agorogiannis, A. Papadimitriou, G. Hadjigeorgiou. Protein misfolding in neurodegenerative diseases. Neuropathol. appl. neurobiol., 2004, 30, 215. https://doi.org/10.1111/j.1365-2990.2004.00558.x DOI: https://doi.org/10.1111/j.1365-2990.2004.00558.x
K. Marciniuk, R. Taschuk, S. Napper. Evidence for prion‐like mechanisms in several neurodegenerative diseases: Potential implications for immunotherapy. J. Immunol. Res., 2013, 2013, 473706. https://doi.org/10.1155/2013/473706 DOI: https://doi.org/10.1155/2013/473706
T. R. Jahn, O. S. Makin, K. L. Morris, K. E. Marshall, P. Tian, P. Sikorski, L. C. Serpell. The common architecture of cross-β amyloid. J. Mol. Biol., 2010, 395, 717 https://doi.org/10.1016/j.jmb.2009.09.039 DOI: https://doi.org/10.1016/j.jmb.2009.09.039
S. Tzotzos, A. J. Doig. Amyloidogenic sequences in native protein structures. Protein Sci., 2010, 19, 327. https://doi.org/10.1002/pro.314 DOI: https://doi.org/10.1002/pro.314
L. C. Serpell. Alzheimer’s amyloid fibrils: structure and assembly. Biochim. Biophys. Acta Mol. Basis Dis., 2000, 1502, 16. https://doi.org/10.1073/pnas.95.11.6448 DOI: https://doi.org/10.1016/S0925-4439(00)00029-6
S. Kumar, J. B. Udgaonkar. Mechanisms of amyloid fibril formation by proteins. Curr. Sci., 2010, 639.
T. Sinnige. Molecular mechanisms of amyloid formation in living systems. Chem. Sci., 2022, 13, 7080. https://doi.org/10.1039/D2SC01278B DOI: https://doi.org/10.1039/D2SC01278B
C. M. Dobson. Protein folding and misfolding. Nature 2003, 426, 884. https://doi.org/10.1038/nature02261 DOI: https://doi.org/10.1038/nature02261
W. E. Balch, R. I. Morimoto, A. Dillin, J. W. Kelly. Adapting proteostasis for disease intervention. Science 2008, 319, 916. https://doi.org/10.1126/science.1141448 DOI: https://doi.org/10.1126/science.1141448
A. K. Buell, C. M. Dobson, T. P. Knowles. The physical chemistry of the amyloid phenomenon: thermodynamics and kinetics of filamentous protein aggregation. Essays Biochem., 2014, 56, 11. https://doi.org/10.1042/bse0560011 DOI: https://doi.org/10.1042/bse0560011
S. I. Cohen, S. Linse, L. M. Luheshi, E. Hellstrand, D. A. White, L. Rajah, D. E. Otzen, M. Vendruscolo, C. M. Dobson, T. P. Knowles. Proliferation of amyloid-β42 aggregates occurs through a secondary nucleation mechanism. Proc Natl Acad Sci U.S.A., 2013, 110, 9758. https://doi.org/10.1073/pnas.1218402110 DOI: https://doi.org/10.1073/pnas.1218402110
F. Bemporad, G. Calloni, S. Campioni, G. Plakoutsi, N. Taddei, F. Chiti. Sequence and structural determinants of amyloid fibril formation. Acc. Chem. Res., 2006, 39, 620. https://doi.org/10.1021/ar050067x DOI: https://doi.org/10.1021/ar050067x
G. G. Tartaglia, M. Vendruscolo. The Zyggregator method for predicting protein aggregation propensities. Chem. Soc. Rev., 2008, 37, 1395. https://doi.org/10.1039/B706784B DOI: https://doi.org/10.1039/b706784b
R. G. Roy, P. K. Mandal, J. C. Maroon. Oxidative stress occurs prior to amyloid Aβ plaque formation and tau phosphorylation in Alzheimer’s disease: Role of glutathione and metal ions. ACS Chem. Neurosci., 2023, 14, 2944. https://doi.org/10.1021/acschemneuro.3c00486 DOI: https://doi.org/10.1021/acschemneuro.3c00486
H.-g. Lee, G. Perry, P. I. Moreira, M. R. Garrett, Q. Liu, X. Zhu, A. Takeda, A. Nunomura, M. A. Smith. Tau phosphorylation in Alzheimer's disease: pathogen or protector? Trends Mol. Med., 2005, 11, 164. https://doi.org/10.1016/j.molmed.2005.02.008 DOI: https://doi.org/10.1016/j.molmed.2005.02.008
A. P. Minton. Influence of macromolecular crowding upon the stability and state of association of proteins: predictions and observations. J. Pharm. Sci., 2005, 94, 1668. https://doi.org/10.1002/jps.20417 DOI: https://doi.org/10.1002/jps.20417
R. J. Ellis, A. P. Minton. Protein aggregation in crowded environments. Biological Chemistry 2006, 387, 485. https://doi.org/10.1515/bc.2006.064 DOI: https://doi.org/10.1515/BC.2006.064
T. R. Jahn, S. E. Radford. Folding versus aggregation: polypeptide conformations on competing pathways. Arch. Biochem. Biophys., 2008, 469, 100. https://doi.org/10.1016/j.abb.2007.05.015 DOI: https://doi.org/10.1016/j.abb.2007.05.015
C. Pintado-Grima, O. Bárcenas, A. Bartolomé-Nafría, M. Fornt-Suñé, V. Iglesias, J. Garcia-Pardo, S. Ventura. A review of fifteen years developing computational tools to study protein aggregation. Biophysica 2023, 3, 1. https://doi.org/10.3390/biophysica3010001 DOI: https://doi.org/10.3390/biophysica3010001
M. P. Lambert, A. Barlow, B. A. Chromy, C. Edwards, R. Freed, M. Liosatos, T. Morgan, I. Rozovsky, B. Trommer, K. L. Viola. Diffusible, nonfibrillar ligands derived from Aβ1–42 are potent central nervous system neurotoxins. Proc. Natl. Acad. Sci. U.S.A., 1998, 95, 6448. https://doi.org/10.1073/pnas.95.11.6448 DOI: https://doi.org/10.1073/pnas.95.11.6448
I. Benilova, E. Karran, B. De Strooper. The toxic Aβ oligomer and Alzheimer's disease: an emperor in need of clothes. Nat. Neurosci., 2012, 15, 349. https://doi.org/10.1038/nn.3028 DOI: https://doi.org/10.1038/nn.3028
R. I. Horne, S. E. Sandler, M. Vendruscolo, U. F. Keyser. Detection of protein oligomers with nanopores. Nat. Rev. Chem., 2025, 9, 224. https://doi.org/10.1038/s41570-025-00694-7 DOI: https://doi.org/10.1038/s41570-025-00694-7
H. Chakraborty, A. Chattopadhyay. Excitements and challenges in GPCR oligomerization: molecular insight from FRET. ACS Chem. Neurosci., 2015, 6, 199. https://doi.org/10.1021/cn500231d DOI: https://doi.org/10.1021/cn500231d
R. Tycko. Molecular structure of amyloid fibrils: insights from solid-state NMR. Q. Rev. Biophys., 2006, 39, 1. https://doi.org/10.1017/S0033583506004173 DOI: https://doi.org/10.1017/S0033583506004173
R. Tycko. Solid-state NMR studies of amyloid fibril structure. Annu. Rev. Phys. Chem., 2011, 62, 279. https://doi.org/10.1146/annurev-physchem-032210-103539 DOI: https://doi.org/10.1146/annurev-physchem-032210-103539
A. W. Fitzpatrick, B. Falcon, S. He, A. G. Murzin, G. Murshudov, H. J. Garringer, R. A. Crowther, B. Ghetti, M. Goedert, S. H. Scheres. Cryo-EM structures of tau filaments from Alzheimer’s disease. Nature 2017, 547, 185. https://doi.org/10.1038/nature23002 DOI: https://doi.org/10.1038/nature23002
A. W. Fitzpatrick, H. R. Saibil. Cryo-EM of amyloid fibrils and cellular aggregates. Curr. Opin. Struct. Biol., 2019, 58, 34. https://doi.org/10.1016/j.sbi.2019.05.003 DOI: https://doi.org/10.1016/j.sbi.2019.05.003
A. -M. Chiorcea-Paquim, W. B. S. Machini, A. M. Oliveira-Brett. Amyloid–ß peptides interaction with curcumin: AFM and electrochemical characterisation. Electrochim. Acta 2025, 525, 146160. https://doi.org/10.1016/j.electacta.2025.146160 DOI: https://doi.org/10.1016/j.electacta.2025.146160
R. Khurana, C. Ionescu-Zanetti, M. Pope, J. Li, L. Nielson, M. Ramírez-Alvarado, L. Regan, A. L. Fink, S. A. Carter. A general model for amyloid fibril assembly based on morphological studies using atomic force microscopy. Biophys. J., 2003, 85, 1135. https://doi.org/10.1016/S0006-3495(03)74550-0 DOI: https://doi.org/10.1016/S0006-3495(03)74550-0
A. J. Dear, T. C. Michaels, G. Meisl, D. Klenerman, S. Wu, S. Perrett, S. Linse, C. M. Dobson, T. P. Knowles. Kinetic diversity of amyloid oligomers. Proc. Natl. Acad. Sci. U.S.A., 2020, 117, 12087. https://doi.org/10.1073/pnas.1922267117 DOI: https://doi.org/10.1073/pnas.1922267117
S. Bhattacharya, L. Xu, D. Thompson. Revisiting the earliest signatures of amyloidogenesis: Roadmaps emerging from computational modeling and experiment. Wiley Interdiscip. Rev. Comput. Mol. Sci., 2018, 8, e1359. https://doi.org/10.1002/wcms.1359 DOI: https://doi.org/10.1002/wcms.1359
Z. Faidon Brotzakis, T. Lohr, S. Truong, S. Hoff, M. Bonomi, M. Vendruscolo. Determination of the structure and dynamics of the fuzzy coat of an amyloid fibril of IAPP using cryo-electron microscopy. Biochem., 2023, 62, 2407. https://doi.org/10.1021/acs.biochem.3c00010 DOI: https://doi.org/10.1021/acs.biochem.3c00010
M. Schäffler, D. J. Wales, B. Strodel. The energy landscape of Aβ 42: a funnel to disorder for the monomer becomes a folding funnel for self-assembly. Chem. Commun., 2024, 60, 13574. https://doi.org/10.1039/D4CC02856B DOI: https://doi.org/10.1039/D4CC02856B
A. V. Kuznetsov. Numerical and analytical simulation of the growth of amyloid-β plaques. J. Biomech. Eng., 2024, 146, 061004. https://doi.org/10.1115/1.4064969 DOI: https://doi.org/10.1115/1.4064969
K. Khalili, F. Farzam, B. Dabirmanesh, K. Khajeh. Prediction of protein aggregation. Prog. Mol. Biol. Transl. Sci., 2024, 206, 229. https://doi.org/10.1016/bs.pmbts.2024.03.005 DOI: https://doi.org/10.1016/bs.pmbts.2024.03.005
V. Cima, A. Kunka, E. Grakova, J. Planas-Iglesias, M. Havlasek, M. Subramanian, M. Beloch, M. Marek, K. Slaninova, J. Damborsky. Prediction of aggregation prone regions in proteins using deep neural networks and their suppression by computational design. bioRxiv 2024, 2024.03. 06.583680. https://doi.org/10.1101/2024.03.06.583680 DOI: https://doi.org/10.1101/2024.03.06.583680
B. Banik (ed). Green Approaches in Medicinal Chemistry for Sustainable Drug Design: Applications, 2nd ed.,Vol. 1, Elsevier, 2024. DOI: https://doi.org/10.1016/B978-0-443-16166-7.00029-3
B. Rebecca, K. P. M. Kumar, S. Padmini, B. K. Srivastava, S. Halder, S. Boopathi, in Handbook of Research on AI and ML for Intelligent Machines and Systems, IGI Global Scientific Publishing, 2024, pp. 348-373. DOI: https://doi.org/10.4018/978-1-6684-9999-3.ch014
C. Soto, L. Estrada. Amyloid inhibitors and β-sheet breakers. In: J. R. Harris, F. Fahrenholz (eds), Alzheimer’s Disease. Subcellular Biochemistry, vol 38. Springer, Boston, MA. 2005, 351 https://doi.org/10.1007/0-387-23226-5_18 DOI: https://doi.org/10.1007/0-387-23226-5_18
Y. -L. Han, C. Li, H. -H. Yin, Y. He, Y. -X. Guan. In Silico Screening of Small Molecule Inhibitors for Amyloid-β Aggregation. J. Chem. Inf. Model., 2025, 65, 6238. https://doi.org/10.1021/acs.jcim.5c00831 DOI: https://doi.org/10.1021/acs.jcim.5c00831
T. Qureshi, S. E. Desale, H. Chidamabram, S. Chinnathambi, in Multi-Factorial Approach as a Therapeutic Strategy for the Management of Alzheimer’s Disease, Springer, 2025, pp. 113-138. DOI: https://doi.org/10.1007/978-981-96-0259-9_6
M. A. Alwaili, F. F. A. Alkhayl, H. A. Rudayni, A. A. Allam, N. G. Altoom, A. M. Lamsabhi, E. M. Kamel. Unraveling molecular mechanisms of β-glucuronidase inhibition by flavonoids from Centaurea scoparia: integrated in silico and in vitro insights. New J. Chem., 2024, 48, 14236. https://doi.org/10.1039/D4NJ02393E DOI: https://doi.org/10.1039/D4NJ02393E
W. Li, R. Cao, J. Cao, J. Zhang, X. Yang, X. Liu, L. Sun. Proteins make tea polyphenol EGCG not always develop satisfactory α-glucosidase inhibition: The influences of three proteins on α-glucosidase inhibitory activity of EGCG. Food Hydrocoll., 2025, 159, 110674. https://doi.org/10.1016/j.foodhyd.2024.110674 DOI: https://doi.org/10.1016/j.foodhyd.2024.110674
O. D. Warerkar, N. H. Mudliar, M. M. Momin, P. K. Singh. Targeting amyloids with coated nanoparticles: a review on potential combinations of nanoparticles and bio-compatible coatings. Crit. Rev. Ther. Drug Carrier Syst., 2024, 41, 85. https://doi.org/10.1615/critrevtherdrugcarriersyst.2023046209 DOI: https://doi.org/10.1615/CritRevTherDrugCarrierSyst.2023046209
Q. -Q. Wang. Anion recognition-directed supramolecular catalysis with functional macrocycles and molecular cages. Acc. Chem. Res., 2024, 57, 3227. https://doi.org/10.1021/acs.accounts.4c00583 DOI: https://doi.org/10.1021/acs.accounts.4c00583
F. A. Mohammed, T. Xiao, L. Wang, R. B. Elmes. Macrocyclic receptors for anion recognition. Chem. Commun., 2024, 60, 11812. https://doi.org/10.1039/D4CC04521A DOI: https://doi.org/10.1039/D4CC04521A
P. Ghosh, A. Bera, P. Bhadury, P. De. From small molecules to synthesized polymers: potential role in combating amyloidogenic disorders. ACS Chem. Neurosci., 2021, 12, 1737. https://doi.org/10.1021/acschemneuro.1c00104 DOI: https://doi.org/10.1021/acschemneuro.1c00104
M. Holubová, P. Štěpánek, M. Hrubý. Polymer materials as promoters/inhibitors of amyloid fibril formation. Colloid Polym. Sci., 2021, 299, 343. https://doi.org/10.1007/s00396-020-04710-8 DOI: https://doi.org/10.1007/s00396-020-04710-8
Y. Niu, P. Lin. Advances of computer-aided drug design (CADD) in the development of anti-Azheimer’s-disease drugs. Drug Discov. Today 2023, 28, 103665. https://doi.org/10.1016/j.drudis.2023.103665 DOI: https://doi.org/10.1016/j.drudis.2023.103665
B. Devanath, S. Deka, D. Das, H. R. Bhat, N. R. Chanu, K. Bhattacharya, in Computational and Experimental Studies in Alzheimer's Disease, CRC Press, 2024, pp. 53-70. DOI: https://doi.org/10.1201/9781003412069-4
F. Chiti, C. M. Dobson. Protein misfolding, amyloid formation, and human disease: a summary of progress over the last decade. Annu. Rev. Biochem., 2017, 86, 27. https://doi.org/10.1146/annurev-biochem-061516-045115 DOI: https://doi.org/10.1146/annurev-biochem-061516-045115
D. Eisenberg, M. Jucker. The amyloid state of proteins in human diseases. Cell 2012, 148, 1188. https://doi.org/10.1038/nn.3028 DOI: https://doi.org/10.1016/j.cell.2012.02.022
S. S. Mamsa, B. P. Meloni. Arginine and arginine-rich peptides as modulators of protein aggregation and cytotoxicity associated with Alzheimer’s disease. Front. Mol. Neurosci., 2021, 14, 759729. https://doi.org/10.3389/fnmol.2021.759729 DOI: https://doi.org/10.3389/fnmol.2021.759729
M. Matveyenka, S. Rizevsky, D. Kurouski. Amyloid aggregates exert cell toxicity causing irreversible damages in the endoplasmic reticulum. Biochim. Biophys. Acta Mol. Basis Dis., 2022, 1868, 166485. https://doi.org/10.1016/j.bbadis.2022.166485 DOI: https://doi.org/10.1016/j.bbadis.2022.166485
T. C. Michaels, D. Qian, A. Šarić, M. Vendruscolo, S. Linse, T. P. Knowles. Amyloid formation as a protein phase transition. Nat. Rev. Phys., 2023, 5, 379. https://doi.org/10.1038/s42254-023-00598-9 DOI: https://doi.org/10.1038/s42254-023-00598-9
R. Cascella, A. Bigi, N. Cremades, C. Cecchi. Effects of oligomer toxicity, fibril toxicity and fibril spreading in synucleinopathies. Cell. Mol. Life Sci., 2022, 79, 174. https://doi.org/10.1007/s00018-022-04166-9 DOI: https://doi.org/10.1007/s00018-022-04166-9
A. Nutini. Amyloid oligomers and their membrane toxicity-A perspective study. Prog. Biophys. Mol. Biol., 2024, 187, 9. https://doi.org/10.1016/j.pbiomolbio.2024.01.002 DOI: https://doi.org/10.1016/j.pbiomolbio.2024.01.002
Z. Du, M. Li, J. Ren, X. Qu. Current strategies for modulating Aβ aggregation with multifunctional agents. Acc. Chem. Res., 2021, 54, 2172. https://doi.org/10.1021/acs.accounts.1c00055 DOI: https://doi.org/10.1021/acs.accounts.1c00055
F. U. Hartl, A. Bracher, M. Hayer-Hartl. Molecular chaperones in protein folding and proteostasis. Nature 2011, 475, 324. https://doi.org/10.1038/nature10317 DOI: https://doi.org/10.1038/nature10317
S. Zampar, S. E. Di Gregorio, G. Grimmer, J. C. Watts, M. Ingelsson. “Prion-like” seeding and propagation of oligomeric protein assemblies in neurodegenerative disorders. Front. Neurosci., 2024, 18, 1436262. https://doi.org/10.3389/fnins.2024.1436262 DOI: https://doi.org/10.3389/fnins.2024.1436262
Y. Baek, M. Lee. Exploring the complexity of amyloid-beta fibrils: structural polymorphisms and molecular interactions. Biochem. Soc. Trans., 2024, 52, 1631. https://doi.org/10.1042/BST20230854 DOI: https://doi.org/10.1042/BST20230854
S. Lövestam, D. Li, J. L. Wagstaff, A. Kotecha, D. Kimanius, S. H. McLaughlin, A. G. Murzin, S. M. Freund, M. Goedert, S. H. Scheres. Disease-specific tau filaments assemble via polymorphic intermediates. Nature 2024, 625, 119. https://doi.org/10.1038/s41586-023-06788-w DOI: https://doi.org/10.1038/s41586-023-06788-w
W. Puławski, R. Dec, W. Dzwolak. Clues to the Design of Aggregation-Resistant Insulin from proline scanning of highly Amyloidogenic peptides derived from the N-terminal segment of the A-chain. Mol. Pharm., 2024, 21, 2025. https://doi.org/10.1021/acs.molpharmaceut.4c00077 DOI: https://doi.org/10.1021/acs.molpharmaceut.4c00077
W. Puławski, A. Koliński, M. Koliński. Multiscale modeling of protofilament structures: A case study on insulin amyloid aggregates. Int. J. Biol. Macromol., 2025, 285, 138382. https://doi.org/10.1016/j.ijbiomac.2024.138382 DOI: https://doi.org/10.1016/j.ijbiomac.2024.138382
Q. Xu, Y. Ma, Y. Sun, D. Li, X. Zhang, C. Liu. Protein amyloid aggregate: Structure and function. Aggregate 2023, 4, e333. https://doi.org/10.1002/agt2.333 DOI: https://doi.org/10.1002/agt2.333
R. Limbocker, N. Cremades, R. Cascella, P. M. Tessier, M. Vendruscolo, F. Chiti. Characterization of Pairs of Toxic and Nontoxic Misfolded Protein Oligomers Elucidates the Structural Determinants of Oligomer Toxicity in Protein Misfolding Diseases. Acc. Chem. Res., 2023, 56, 1395. https://doi.org/10.1021/acs.accounts.3c00045 DOI: https://doi.org/10.1021/acs.accounts.3c00045
A. González Díaz, R. Cataldi, B. Mannini, M. Vendruscolo. Preparation and characterization of zn (ii)-stabilized Aβ42 oligomers. ACS Chem. Neurosci., 2024, 15, 2586. https://doi.org/10.1021/acschemneuro.4c00084 DOI: https://doi.org/10.1021/acschemneuro.4c00084
S. Chia, R. L. Cataldi, F. S. Ruggeri, R. Limbocker, I. Condado-Morales, K. Pisani, A. Possenti, S. Linse, T. P. Knowles, J. Habchi. A relationship between the structures and neurotoxic effects of Aβ oligomers stabilized by different metal ions. ACS Chem. Neurosci., 2024, 15, 1125 https://doi.org/10.1021/acschemneuro.3c00718 DOI: https://doi.org/10.1021/acschemneuro.3c00718
R. Zou, H. Ågren. Molecular Insight into Amyloid Fibril-Templated Aggregation of Biomarkers. ACS Chem. Neurosci., 2025, 16, 2076. https://doi.org/10.1021/acschemneuro.5c00103 DOI: https://doi.org/10.1021/acschemneuro.5c00103
R. Duran-Romaña, J. Schymkowitz, F. Rousseau, N. Louros. Energetic profiling reveals thermodynamic principles underlying amyloid fibril maturation. Nat. Commun., 2025, 16, 10081. https://doi.org/10.1038/s41467-025-65035-0 DOI: https://doi.org/10.1038/s41467-025-65035-0
E. Duran-Meza, R. Araya-Secchi, P. Romero-Hasler, E. A. Soto-Bustamante, V. Castro-Fernandez, C. Castillo-Caceres, O. Monasterio, R. Diaz-Espinoza. Metal Ions Can Modulate the Self-Assembly and Activity of Catalytic Peptide Amyloids. Langmuir 2024, 40, 6094. https://doi.org/10.1021/acs.langmuir.3c02983 DOI: https://doi.org/10.1021/acs.langmuir.3c02983
Y. Wang, Y. Tang, M. Zhao, B. Yan, Y. Zhang, X. Gao, J. Fan, M. Wang. Transition metal ion-catalyzed assembly of oat peptides into amyloid fibrils: Feasibility, physicochemical properties, and chitosan film reinforcement. Food Hydrocoll., 2025, 169, 111628. https://doi.org/10.1016/j.foodhyd.2025.111628 DOI: https://doi.org/10.1016/j.foodhyd.2025.111628
A. P. de Oliveira, D. Baghel, B. Holcombe, W. Chase, T. Ward, S.-H. J. Wang, A. Ghosh. Lipid mediated formation of antiparallel aggregates in cerebral amyloid angiopathy. Acta Neuropathol., 2025, 150, 3. https://doi.org/10.1007/s00401-025-02911-5 DOI: https://doi.org/10.1007/s00401-025-02911-5
S. Efimova, O. Ostroumova. Mechanisms of lipid-mediated regulation of the pore-forming activity of antimicrobial agents: studies on planar lipid bilayers. Biochem. Moscow Suppl. Ser. A 2024, 18, 257. https://doi.org/10.1134/S1990747824700247 DOI: https://doi.org/10.1134/S1990747824700247
R. M. Murphy. Kinetics of amyloid formation and membrane interaction with amyloidogenic proteins. Biochim. Biophys. Acta Biomembr., 2007, 1768, 1923. https://doi.org/10.1016/j.bbamem.2006.12.014 DOI: https://doi.org/10.1016/j.bbamem.2006.12.014
A. Kamada, Z. Toprakcioglu, T. P. Knowles. Kinetic analysis reveals the role of secondary nucleation in regenerated silk fibroin self-assembly. Biomacromolecules 2023, 24, 1709. https://doi.org/10.1021/acs.biomac.2c01479 DOI: https://doi.org/10.1021/acs.biomac.2c01479
J. Wei, G. Meisl, A. J. Dear, T. C. Michaels, T. P. Knowles. Kinetics of Amyloid Oligomer Formation. Annu. Rev. Biophys., 2025, 54, 185. https://doi.org/10.1146/annurev-biophys-080124-122953 DOI: https://doi.org/10.1146/annurev-biophys-080124-122953
C. Tschöpe, A. Elsanhoury, A. V. Kristen. Transthyretin Amyloid Cardiomyopathy—2025 Update: Current Diagnostic Approaches and Emerging Therapeutic Options. J. Clin. Med., 2025, 14, 4785. https://doi.org/10.3390/jcm14134785 DOI: https://doi.org/10.3390/jcm14134785
A. Lalario, R. Saro, G. Sinagra, M. Merlo, A. Porcari. Clinical Use of Biomarkers in Cardiac Amyloidosis. Heart Failure Clinic 2024, 20, 283. https://doi.org/10.1016/j.hfc.2024.03.001 DOI: https://doi.org/10.1016/j.hfc.2024.03.001
A. Albulushi, J. Al Buraiki, G. Aly, Y. Al-Wahshi, A. Jahangirifard. Role of biomarkers in early diagnosis and prognosis of cardiac amyloidosis: A systematic review and meta-analysis. Curr. Probl. Cardiol., 2025, 50, 102883. https://doi.org/10.1016/j.cpcardiol.2024.102883 DOI: https://doi.org/10.1016/j.cpcardiol.2024.102883
N. Vilor-Tejedor, P. Genius, B. Rodríguez-Fernández, C. Minguillón, I. Sadeghi, A. González-Escalante, M. Crous-Bou, M. Suárez-Calvet, O. Grau-Rivera, A. Brugulat-Serrat, G. Sánchez-Benavides, M. Esteller, K. Fauria, J. L. Molinuevo, A. Navarro, J. D. Gispert, f. t. A. s. D. N. Initiative, f. t. A. study. Genetic characterization of the ALFA study: Uncovering genetic profiles in the Alzheimer's continuum. Alzheimer's & Dementia 2024, 20, 1703. https://doi.org/10.1002/alz.13537 DOI: https://doi.org/10.1002/alz.13537
C. Alarcón-Vila, L. Hurtado-Navarro, S. V. Mateo, A. Peñín-Franch, C. M. Martínez, C. Molina-López, M. C. Baños, A. I. Gómez, J. Gómez-Román, A. Baroja-Mazo. The inflammasome adaptor protein ASC promotes amyloid deposition in cryopyrin-associated periodic syndromes. EMBO Mol. Med., 2025, 17, 41. https://doi.org/10.1038/s44321-024-00176-1 DOI: https://doi.org/10.1038/s44321-024-00176-1
Y. Tao, W. Xia, Q. Zhao, H. Xiang, C. Han, S. Zhang, W. Gu, W. Tang, Y. Li, L. Tan. Structural mechanism for specific binding of chemical compounds to amyloid fibrils. Nat. Chem. Biol., 2023, 19, 1235. https://doi.org/10.1038/s41589-023-01370-x DOI: https://doi.org/10.1038/s41589-023-01370-x
T. Tripathi, V. K. Dubey (Eds), Advances in protein molecular and structural biology methods, Academic Press, 2022.
H. Fatafta, M. Khaled, B. Kav, O. O. Olubiyi, B. Strodel. A brief history of amyloid aggregation simulations. Wiley Interdiscip. Rev. Comput. Mol. Sci., 2024, 14, e1703. https://doi.org/10.1002/wcms.1703 DOI: https://doi.org/10.1002/wcms.1703
R. Pandey, B. Urbanc. Oligomer Formation by Physiologically Relevant C-Terminal Isoforms of Amyloid β-Protein. Biomolecules 2024, 14, 774. https://doi.org/10.3390/biom14070774 DOI: https://doi.org/10.3390/biom14070774
S. Bosio, F. Falchi, C. Rauzi, L. Bellucci. Morphological inhibitors of aggregation-prone amyloid-β conformers: A computational exploration. Comput. Biol. Med., 2025, 196, 110545. https://doi.org/10.1016/j.compbiomed.2025.110545 DOI: https://doi.org/10.1016/j.compbiomed.2025.110545
V. Iglesias, J. Chilimoniuk, C. Pintado-Grima, O. Bárcenas, S. Ventura, M. Burdukiewicz. Aggregating amyloid resources: A comprehensive review of databases on amyloid-like aggregation. Comput. Struct. Biotechnol. J., 2024, 23, 4011. https://doi.org/10.1016/j.csbj.2024.10.047 DOI: https://doi.org/10.1016/j.csbj.2024.10.047
M. G. Costa, M. Gur, J. M. Krieger, I. Bahar. Computational biophysics meets cryo‐EM revolution in the search for the functional dynamics of biomolecular systems. Wiley Interdiscip. Rev. Comput. Mol. Sci., 2024, 14, e1689. https://doi.org/10.1002/wcms.1689 DOI: https://doi.org/10.1002/wcms.1689
D. Ghosh, A. Biswas, M. Radhakrishna. Advanced computational approaches to understand protein aggregation. Biophys. rev., 2024, 5, 021302. https://doi.org/10.1063/5.0180691 DOI: https://doi.org/10.1063/5.0180691
A. Morriss-Andrews, J. -E. Shea. Computational Studies of Protein Aggregation: Methods and Applications. Annu. Rev. Phys. Chem., 2015, 66, 643. https://doi.org/10.1146/annurev-physchem-040513-103738 DOI: https://doi.org/10.1146/annurev-physchem-040513-103738
A. Arefeen, S. Singh, C. Razavi, H. Ghasemzadeh, S. Dev. Assessing the quality of reporting in artificial intelligence/machine learning research for cardiac amyloidosis. JAMIA open 2025, 8, ooaf104. https://doi.org/10.1093/jamiaopen/ooaf104 DOI: https://doi.org/10.1093/jamiaopen/ooaf104
A. Barison, D. Tomasoni, A. Filippeschi, M. G. Bellicini, C. A. Avizzano, M. Metra, M. Grogan, in Cardiac Amyloidosis: Diagnosis and Treatment, Springer, 2024, pp. 233-243. DOI: https://doi.org/10.1007/978-3-031-51757-0_19
C. Alfano, Y. Fichou, K. Huber, M. Weiss, E. Spruijt, S. Ebbinghaus, G. De Luca, M. A. Morando, V. Vetri, P. A. Temussi. Molecular crowding: the history and development of a scientific paradigm. Chem. Rev., 2024, 124, 3186. https://doi.org/10.1021/acs.chemrev.3c00615 DOI: https://doi.org/10.1021/acs.chemrev.3c00615
Y. Levites, E. B. Dammer, Y. Ran, W. Tsering, D. Duong, M. Abreha, J. Gadhavi, K. Lolo, J. Trejo-Lopez, J. Phillips. Integrative proteomics identifies a conserved Aβ amyloid responsome, novel plaque proteins, and pathology modifiers in Alzheimer’s disease. Cell Rep. Med., 2024, 5, 101669. https://doi.org/10.1016/j.xcrm.2024.101669 DOI: https://doi.org/10.1016/j.xcrm.2024.101669
A. -L. Mahul-Mellier, M. F. Altay, N. Maharjan, N. Ait-Bouziad, A. Chiki, S. Jagannath, G. Limorenko, S. Novello, J. Ricci, S. Vingill. Dissecting the differential role of C-terminal truncations in the regulation of aSyn pathology formation and the biogenesis of Lewy bodies. bioRxiv 2024, 2024.11. 29.625993. https://doi.org/10.1101/2024.11.29.625993 DOI: https://doi.org/10.1101/2024.11.29.625993
J. Kardos, M. P. Nyiri, É. Moussong, F. Wien, T. Molnár, N. Murvai, V. Tóth, H. Vadászi, J. Kun, F. Jamme. Guide to the structural characterization of protein aggregates and amyloid fibrils by CD spectroscopy. Protein Sci. 2025, 34, e70066. https://doi.org/10.1002/pro.70066 DOI: https://doi.org/10.1002/pro.70066
Đ. Tintor, K. Ninković, J. Milošević, N. Đ. Polović. Gaining insight into protein structure via ATR-FTIR spectroscopy. Vib. Spectrosc., 2024, 134, 103726. https://doi.org/10.1016/j.vibspec.2024.103726 DOI: https://doi.org/10.1016/j.vibspec.2024.103726
M. F. Pignataro, M. G. Herrera, V. I. Dodero. Evaluation of Peptide/Protein Self-Assembly and Aggregation by Spectroscopic Methods. Molecules 2020, 25, 4854. https://doi.org/10.3390/molecules25204854 DOI: https://doi.org/10.3390/molecules25204854
Z. Qin, Y. Sun, B. Jia, D. Wang, Y. Ma, G. Ma. Kinetic mechanism of thioflavin T binding onto the amyloid fibril of hen egg white lysozyme. Langmuir 2017, 33, 5398. https://doi.org/10.1021/acs.langmuir.7b00221 DOI: https://doi.org/10.1021/acs.langmuir.7b00221
E. Dahal, M. Choi, N. Alam, A. A. Bhirde, S. L. Beaucage, A. Badano. Structural evaluation of an amyloid fibril model using small-angle x-ray scattering. Phys. Biol., 2017, 14, 046001. https://doi.org/10.1088/1478-3975/aa776a DOI: https://doi.org/10.1088/1478-3975/aa776a
H. Xie, C. Guo. Albumin alters the conformational ensemble of amyloid-β by promiscuous interactions: Implications for amyloid inhibition. Front. Mol. Biosci., 2021, 7, 629520. https://doi.org/10.3389/fmolb.2020.629520 DOI: https://doi.org/10.3389/fmolb.2020.629520
H. Hampel, J. Hardy, K. Blennow, C. Chen, G. Perry, S. H. Kim, V. L. Villemagne, P. Aisen, M. Vendruscolo, T. Iwatsubo, C. L. Masters, M. Cho, L. Lannfelt, J. L. Cummings, A. Vergallo. The Amyloid-β Pathway in Alzheimer’s Disease. Mol. Psychiatry 2021, 26, 5481. https://doi.org/10.1038/s41380-021-01249-0 DOI: https://doi.org/10.1038/s41380-021-01249-0
D. J. Selkoe, J. Hardy. The amyloid hypothesis of Alzheimer's disease at 25 years. EMBO molecular medicine 2016, 8, 595. https://doi.org/10.15252/emmm.201606210 DOI: https://doi.org/10.15252/emmm.201606210
C. Behl. In 2024, the amyloid-cascade-hypothesis still remains a working hypothesis, no less but certainly no more. Front. Aging Neurosci., 2024, 16, 1459224. https://doi.org/10.3389/fnagi.2024.1459224 DOI: https://doi.org/10.3389/fnagi.2024.1459224
M. Lee, J. Yoon, S. Shin. Computational Study on Structure and Aggregation Pathway of Aβ42 Amyloid Protofibril. J. Phys. Chem. B 2019, 123, 7859. https://doi.org/10.1021/acs.jpcb.9b07195 DOI: https://doi.org/10.1021/acs.jpcb.9b07195
M. Zganec, E. Zerovnik, B. Urbanc. Assembly of stefin B into polymorphic oligomers probed by discrete molecular dynamics. J. Chem. Theory Comput., 2015, 11, 2355. https://doi.org/10.1021/acs.jctc.5b00067 DOI: https://doi.org/10.1021/acs.jctc.5b00067
C. Camilloni, M. Vendruscolo. Statistical mechanics of the denatured state of a protein using replica-averaged metadynamics. J. Am. Chem. Soc., 2014, 136, 8982. https://doi.org/10.1021/ja5027584 DOI: https://doi.org/10.1021/ja5027584
N. Mattsson, S. Palmqvist, E. Stomrud, J. Vogel, O. Hansson. Staging β-Amyloid Pathology With Amyloid Positron Emission Tomography. JAMA Neurol., 2019, 76, 1319. https://doi.org/10.1001/jamaneurol.2019.2214 DOI: https://doi.org/10.1001/jamaneurol.2019.2214
M. Cecchini, F. Rao, M. Seeber, A. Caflisch. Replica exchange molecular dynamics simulations of amyloid peptide aggregation. J. Chem. Phys., 2004, 121, 10748. https://doi.org/10.1063/1.1809588 DOI: https://doi.org/10.1063/1.1809588
X. Li, Z. Yang, Y. Chen, S. Zhang, G. Wei, L. Zhang. Dissecting the molecular mechanisms of the co-aggregation of Aβ40 and Aβ42 peptides: A REMD simulation study. J. Phys. Chem. B 2023, 127, 4050. https://doi.org/10.1021/acs.jpcb.3c01078 DOI: https://doi.org/10.1021/acs.jpcb.3c01078
S. Devkota, R. Zhou, V. Nagarajan, M. Maesako, H. Do, A. Noorani, C. Overmeyer, S. Bhattarai, J. T. Douglas, A. Saraf. Familial Alzheimer mutations stabilize synaptotoxic γ-secretase-substrate complexes. Cell Rep., 2024, 43, 113761. https://doi.org/10.1016/j.celrep.2024.113761 DOI: https://doi.org/10.1016/j.celrep.2024.113761
S. Liu, C. Wang, B. Zhang. Toward Predictive Coarse-Grained Simulations of Biomolecular Condensates. Biochem., 2025, 64, 1750. https://doi.org/10.1021/acs.biochem.4c00737 DOI: https://doi.org/10.1021/acs.biochem.4c00737
D. Chakraborty, J. E. Straub, D. Thirumalai. Energy landscapes of Aβ monomers are sculpted in accordance with Ostwald’s rule of stages. Sci. Adv., 2023, 9, eadd6921. https://doi.org/10.1126/sciadv.add6921 DOI: https://doi.org/10.1126/sciadv.add6921
S. Suresh, A. Singh S, R. Rushendran, C. Vellapandian, B. Prajapati. Alzheimer’s disease: the role of extrinsic factors in its development, an investigation of the environmental enigma. Front. Neurol., 2023, 14, 1303111. https://doi.org/10.3389/fneur.2023.1303111 DOI: https://doi.org/10.3389/fneur.2023.1303111
J. M. Kenyaga, Q. Cheng, W. Qiang. Early stage β-amyloid-membrane interactions modulate lipid dynamics and influence structural interfaces and fibrillation. J. Biol. Chem., 2022, 298, 102491. https://doi.org/10.1016/j.jbc.2022.102491 DOI: https://doi.org/10.1016/j.jbc.2022.102491
M. Goedert, M. G. Spillantini. A tribute to John Q. Trojanowski (1946–2022), neuropathologist extraordinaire. Brain Pathol., 2022, 32, e13066. https://doi.org/10.1111/bpa.13066 DOI: https://doi.org/10.1111/bpa.13066
S. R. Venati, V. N. Uversky. Exploring intrinsic disorder in human synucleins and associated proteins. Int. J. Mol. Sci., 2024, 25, 8399. https://doi.org/10.3390/ijms25158399 DOI: https://doi.org/10.3390/ijms25158399
A. -M. Illig, B. Strodel. Performance of Markov state models and transition networks on characterizing amyloid aggregation pathways from MD data. J. Chem. Theory Comput., 2020, 16, 7825. https://doi.org/10.1021/acs.jctc.0c00727 DOI: https://doi.org/10.1021/acs.jctc.0c00727
C. Privat, S. Madurga, F. Mas, J. Rubio-Martinez. Molecular dynamics simulations of an α-synuclein NAC domain fragment with a ff14IDPSFF IDP-specific force field suggest β-sheet intermediate states of fibrillation. Phys. Chem. Chem. Phys., 2022, 24, 18841. https://doi.org/10.1039/D2CP02042D DOI: https://doi.org/10.1039/D2CP02042D
I. F. Tsigelny, Y. Sharikov, V. L. Kouznetsova, J. P. Greenberg, W. Wrasidlo, C. Overk, T. Gonzalez, M. Trejo, B. Spencer, K. Kosberg. Molecular determinants of α-synuclein mutants’ oligomerization and membrane interactions. ACS Chem. Neurosci., 2015, 6, 403. https://doi.org/10.1021/cn500332w DOI: https://doi.org/10.1021/cn500332w
M. Vidović, M. G. Rikalovic. Alpha-synuclein aggregation pathway in Parkinson’s disease: current status and novel therapeutic approaches. Cells 2022, 11, 1732. https://doi.org/10.3390/cells11111732 DOI: https://doi.org/10.3390/cells11111732
H.-C. Fan, L.-I. Ho, C.-S. Chi, S.-J. Chen, G.-S. Peng, T.-M. Chan, S.-Z. Lin, H.-J. Harn. Polyglutamine (PolyQ) diseases: genetics to treatments. Cell Transplant., 2014, 23, 441. https://doi.org/10.3727/096368914X678454 DOI: https://doi.org/10.3727/096368914X678454
M. D. Costa, P. Maciel. Modifier pathways in polyglutamine (PolyQ) diseases: from genetic screens to drug targets. Cell. Mol. Life Sci., 2022, 79, 274. https://doi.org/10.1007/s00018-022-04280-8 DOI: https://doi.org/10.1007/s00018-022-04280-8
K. M. Ruff, S. J. Khan, R. V. Pappu. A coarse-grained model for polyglutamine aggregation modulated by amphipathic flanking sequences. Biophys. J., 2014, 107, 1226. https://doi.org/10.1016/j.bpj.2014.07.019 DOI: https://doi.org/10.1016/j.bpj.2014.07.019
M. Farag, R. V. Pappu. Physical descriptions of molecular and mesoscale organizations of proteins within condensates and at interfaces. Biophys. J., 2022, 121, 355a. https://doi.org/10.1016/j.bpj.2021.11.960 DOI: https://doi.org/10.1016/j.bpj.2021.11.960
A. Böker, W. Paul. Thermodynamics and conformations of single polyalanine, polyserine, and polyglutamine chains within the PRIME20 model. J. Phys. Chem. B 2022, 126, 7286. https://doi.org/10.1021/acs.jpcb.2c04360 DOI: https://doi.org/10.1021/acs.jpcb.2c04360
J. A. Larsen, J. H. van Gils, S. Ray, M. Dickmanns, S. Wang, A. Sadek, H. Mohammad-Beigi, M. Zanganeh, S. Abeln, A. K. Buell. Probing the effect of the disordered flank regions on amyloid fibril growth and proliferation. RSC Adv. 2025, 15, 20668. https://doi.org/10.1039/D5RA01654A DOI: https://doi.org/10.1039/D5RA01654A
X. Wang, S. Zhang, J. Zhang, Y. Wang, X. Jiang, Y. Tao, D. Li, C. Zhong, C. Liu. Rational design of functional amyloid fibrillar assemblies. Chem. Soc. Rev., 2023, 52, 4603. https://doi.org/10.1039/D2CS00756H DOI: https://doi.org/10.1039/D2CS00756H
H. F. Fisher, R. J. Boys, C. S. Gillespie, C. J. Proctor, A. Golightly. Parameter inference for a stochastic kinetic model of expanded polyglutamine proteins. Biometrics 2022, 78, 1195. https://doi.org/10.1111/biom.13467 DOI: https://doi.org/10.1111/biom.13467
A. Ganne, M. Balasubramaniam, S. Ayyadevara, R. J. Shmookler Reis. Machine-learning analysis of intrinsically disordered proteins identifies key factors that contribute to neurodegeneration-related aggregation. Front. Aging Neurosci., 2022, 14, 938117. https://doi.org/10.3389/fnagi.2022.938117 DOI: https://doi.org/10.3389/fnagi.2022.938117
P. Mier, M. A. Andrade-Navarro. Predicting the involvement of polyQ-and polyA in protein-protein interactions by their amino acid context. Heliyon 2024, 10, e37861. https://doi.org/10.1016/j.heliyon.2024.e37861 DOI: https://doi.org/10.1016/j.heliyon.2024.e37861
E. A. Algharably, E. Di Consiglio, E. Testai, F. Pistollato, H. Mielke, U. Gundert-Remy. In vitro–in vivo extrapolation by physiologically based kinetic modeling: Experience with three case studies and lessons learned. Front. Toxicol., 2022, 4, 885843. https://doi.org/10.3389/ftox.2022.885843 DOI: https://doi.org/10.3389/ftox.2022.885843
B. S. Visser, W. P. Lipiński, E. Spruijt. The role of biomolecular condensates in protein aggregation. Nat. Rev. Chem., 2024, 8, 686. https://doi.org/10.1038/s41570-024-00635-w DOI: https://doi.org/10.1038/s41570-024-00635-w
D. Kumar, G. M. Hasan, A. Islam, M. I. Hassan. Therapeutic targeting of Huntington's disease: Molecular and clinical approaches. Biochem. Biophys. Res. Commun., 2023, 655, 18. https://doi.org/10.1016/j.bbrc.2023.02.075 DOI: https://doi.org/10.1016/j.bbrc.2023.02.075
J. Cummings. Anti-amyloid monoclonal antibodies are transformative treatments that redefine Alzheimer's disease therapeutics. Drugs 2023, 83, 569. https://doi.org/10.1007/s40265-023-01858-9 DOI: https://doi.org/10.1007/s40265-023-01858-9
R. Dobson, K. Patterson, R. Malik, U. Mandal, H. Asif, R. Humphreys, M. Payne, E. O-Charoenrat, L. Huzzey, A. Clare. Eligibility for antiamyloid treatment: preparing for disease-modifying therapies for Alzheimer’s disease. J. Neurol. Neurosurg. Psychiatry 2024, 95, 796. https://doi.org/10.1136/jnnp-2024-333468 DOI: https://doi.org/10.1136/jnnp-2024-333468
C. Castiello, P. Junghanns, A. Mergel, C. Jacob, C. Ducho, S. Valente, D. Rotili, R. Fioravanti, C. Zwergel, A. Mai. GreenMedChem: the challenge in the next decade toward eco-friendly compounds and processes in drug design. Green Chem., 2023, 25, 2109. https://doi.org/10.1039/D2GC03772F DOI: https://doi.org/10.1039/D2GC03772F
J. Becker, C. Manske, S. Randl. Green chemistry and sustainability metrics in the pharmaceutical manufacturing sector. Curr. Opin. Green Sustain. Chem., 2022, 33, 100562. https://doi.org/10.1016/j.cogsc.2021.100562 DOI: https://doi.org/10.1016/j.cogsc.2021.100562
A. Bera, D. Mukhopadhyay, K. Goswami, P. Ghosh, R. De, P. De. Fatty acid-based polymeric micelles to ameliorate amyloidogenic disorders. Biomater. Sci., 2022, 10, 3466. http://dx.doi.org/10.1039/D2BM00359G DOI: https://doi.org/10.1039/D2BM00359G
A. Dey, U. Haldar, T. Rajasekhar, P. Ghosh, R. Faust, P. De. Polyisobutylene-based glycopolymers as potent inhibitors for in vitro insulin aggregation. J. Mater. Chem. B 2022, 10, 9446. http://dx.doi.org/10.1039/D2TB01856J DOI: https://doi.org/10.1039/D2TB01856J
A. Bera, P. Ghosh, S. Barman, S. Bhattacharya, B. Sudhamalla, K. Goswami, P. De. Insulin fibril inhibition using glycopolymeric nanoassemblies. Biomater. Sci., 2023, 11, 3574. http://dx.doi.org/10.1039/D2BM02078E DOI: https://doi.org/10.1039/D2BM02078E
P. H. Nguyen, A. Ramamoorthy, B. R. Sahoo, J. Zheng, P. Faller, J. E. Straub, L. Dominguez, J.-E. Shea, N. V. Dokholyan, A. De Simone, B. Ma, R. Nussinov, S. Najafi, S. T. Ngo, A. Loquet, M. Chiricotto, P. Ganguly, J. McCarty, M. S. Li, C. Hall, Y. Wang, Y. Miller, S. Melchionna, B. Habenstein, S. Timr, J. Chen, B. Hnath, B. Strodel, R. Kayed, S. Lesné, G. Wei, F. Sterpone, A. J. Doig, P. Derreumaux. Amyloid Oligomers: A Joint Experimental/Computational Perspective on Alzheimer’s Disease, Parkinson’s Disease, Type II Diabetes, and Amyotrophic Lateral Sclerosis. Chem. Rev., 2021, 121, 2545. https://doi.org/10.1021/acs.chemrev.0c01122 DOI: https://doi.org/10.1021/acs.chemrev.0c01122
I. Sedov, D. Khaibrakhmanova. Molecular mechanisms of inhibition of protein amyloid fibril formation: Evidence and perspectives based on kinetic models. Int. J. Mol. Sci., 2022, 23, 13428. https://doi.org/10.3390/ijms232113428 DOI: https://doi.org/10.3390/ijms232113428
R. Swetha, A. Sharma, R. Singh, A. Ganeshpurkar, D. Kumar, A. Kumar, S. K. Singh. Combined ligand-based and structure-based design of PDE 9A inhibitors against Alzheimer’s disease. Mol. Divers., 2022, 26, 2877. https://doi.org/10.1007/s11030-022-10504-7 DOI: https://doi.org/10.1007/s11030-022-10504-7
T. S. Chisholm, M. Mackey, C. A. Hunter. Discovery of high-affinity amyloid ligands using a ligand-based virtual screening pipeline. J. Am. Chem. Soc., 2023, 145, 15936. https://doi.org/10.1021/jacs.3c03749 DOI: https://doi.org/10.1021/jacs.3c03749
V. K. Vyas, S. Bhati, S. Patel, M. Ghate. Structure-and ligand-based drug design methods for the modeling of antimalarial agents: A review of updates from 2012 onwards. J. Biomol. Struct. Dyn., 2022, 40, 10481. https://doi.org/10.1080/07391102.2021.1932598 DOI: https://doi.org/10.1080/07391102.2021.1932598
F. K. Zaidi, R. Bhat. Two polyphenols with diverse mechanisms towards amyloidosis: differential modulation of the fibrillation pathway of human lysozyme by curcumin and EGCG. J. Biomol. Struct. Dyn., 2022, 40, 4593. https://doi.org/10.1080/07391102.2020.1860824 DOI: https://doi.org/10.1080/07391102.2020.1860824
M. Ahmed. Targeting aging pathways with natural compounds: a review of curcumin, epigallocatechin gallate, thymoquinone, and resveratrol. Immun Ageing 2025, 22, 28. https://doi.org/10.1186/s12979-025-00522-y DOI: https://doi.org/10.1186/s12979-025-00522-y
T. Kirkman, C. dos Santos Silva, M. Tosin, M. V. Bertacine Dias. How to find a fragment: methods for screening and validation in fragment‐based drug discovery. ChemMedChem 2024, 19, e202400342. https://doi.org/10.1002/cmdc.202400342 DOI: https://doi.org/10.1002/cmdc.202400342
B. W. Chan, N. B. Lynch, W. Tran, J. M. Joyce, G. P. Savage, W. Meutermans, A. P. Montgomery, M. Kassiou. Fragment-based drug discovery for disorders of the central nervous system: designing better drugs piece by piece. Front. Chem., 2024, 12, 1379518. https://doi.org/10.3389/fchem.2024.1379518 DOI: https://doi.org/10.3389/fchem.2024.1379518
A. A. Ashkarran, S. Tadjiki, Z. Lin, K. Hilsen, N. Ghazali, S. Krikor, S. Sharifi, M. Asgari, M. Hotchkin, A. Dorfman. Protein corona composition of gold nanocatalysts. ACS Pharmacol. Transl. Sci., 2024, 7, 1169 https://doi.org/10.1021/acsptsci.4c00028 DOI: https://doi.org/10.1021/acsptsci.4c00028
A. A. Ashkarran, H. Gharibi, S. A. Sadeghi, S. M. Modaresi, Q. Wang, T.-J. Lin, G. Yerima, A. Tamadon, M. Sayadi, M. Jafari. Small molecule modulation of protein corona for deep plasma proteome profiling. Nat. Commun., 2024, 15, 9638. https://doi.org/10.1038/s41467-024-53966-z DOI: https://doi.org/10.1038/s41467-024-53966-z
X. Shao, C. Yan, C. Wang, C. Wang, Y. Cao, Y. Zhou, P. Guan, X. Hu, W. Zhu, S. Ding. Advanced nanomaterials for modulating Alzheimer's related amyloid aggregation. Nanoscale Adv., 2023, 5, 46. https://doi.org/10.1039/D2NA00625A DOI: https://doi.org/10.1039/D2NA00625A
X. Yao, Y. Guan, J. Wang, D. Wang. Cerium oxide nanoparticles modulating the Parkinson's disease conditions: From the alpha synuclein structural point of view and antioxidant properties of cerium oxide nanoparticles. Heliyon 2024, 10, e21789. https://doi.org/10.1016/j.heliyon.2023.e21789 DOI: https://doi.org/10.1016/j.heliyon.2023.e21789
A. Biswas, P. Hivare, R. Solanki, S. Gupta, D. Bhatia. Applications of bionanomaterials in neurodegenerative diseases. Mater. Adv., 2025, 6, 3785. https://doi.org/10.1039/D4MA01215A DOI: https://doi.org/10.1039/D4MA01215A
A. Bera, P. Ghosh, K. Goswami, P. De. Amino Acid-Based Polymer-Coated Silver Nanoparticles as Insulin Fibril Inhibitors. ACS Appl. Nano Mater., 2023, 6, 8705. https://doi.org/10.1021/acsanm.3c01078 DOI: https://doi.org/10.1021/acsanm.3c01078
L. Wang, N. Kang, H. Shi, L. Gao. Carbon nanotubes and carbon nanoonions inhibit the formation of amyloid fibrils from whey protein isolate. Carbon Trends 2025, 20, 100532. https://doi.org/10.1016/j.cartre.2025.100532 DOI: https://doi.org/10.1016/j.cartre.2025.100532
M. Bazi Alahri, A. Jibril Ibrahim, M. Barani, H. Arkaban, S. M. Shadman, S. Salarpour, P. Zarrintaj, J. Jaberi, A. Turki Jalil. Management of brain cancer and neurodegenerative disorders with polymer-based nanoparticles as a biocompatible platform. Molecules 2023, 28, 841. https://doi.org/10.3390/molecules28020841 DOI: https://doi.org/10.3390/molecules28020841
M. Perxés Perich, S. Palma-Florez, C. Solé, S. Goberna-Ferrón, J. Samitier, P. Gómez-Romero, M. Mir, A. Lagunas. Polyoxometalate-Decorated Gold Nanoparticles Inhibit β-Amyloid Aggregation and Cross the Blood–Brain Barrier in a µphysiological Model. Nanomater., 2023, 13, 2697. https://doi.org/10.3390/nano13192697 DOI: https://doi.org/10.3390/nano13192697
D. T. Dang. Molecular approaches to protein dimerization: opportunities for supramolecular chemistry. Front. Chem., 2022, 10, 829312. https://doi.org/10.3389/fchem.2022.829312 DOI: https://doi.org/10.3389/fchem.2022.829312
K. Taş, B. D. Volta, C. Lindner, O. El Bounkari, K. Hille, Y. Tian, X. Puig-Bosch, M. Ballmann, S. Hornung, M. Ortner. Designed peptides as nanomolar cross-amyloid inhibitors acting via supramolecular nanofiber co-assembly. Nat. Commun., 2022, 13, 5004. https://doi.org/10.1038/s41467-022-32688-0 DOI: https://doi.org/10.1038/s41467-022-32688-0
H. Yin, Q. Cheng, D. Bardelang, R. Wang. Challenges and opportunities of functionalized cucurbiturils for biomedical applications. JACS Au 2023, 3, 2356. https://doi.org/10.1021/jacsau.3c00273 DOI: https://doi.org/10.1021/jacsau.3c00273
Y. Huo, J. Hu, Y. Yin, P. Liu, K. Cai, W. Ji. Self‐assembling peptide‐based functional biomaterials. ChemBioChem 2023, 24, e202200582. https://doi.org/10.1002/cbic.202200582 DOI: https://doi.org/10.1002/cbic.202200582
K. Nayak, P. Ghosh, M. E. H. Khan, P. De. Side-chain amino-acid-based polymers: self-assembly and bioapplications. Polym. Int., 2022, 71, 411. https://doi.org/10.1002/pi.6278 DOI: https://doi.org/10.1002/pi.6278
P. Ghosh, A. Kundu, D. Ganguly. From experimental studies to computational approaches: recent trends in designing novel therapeutics for amyloidogenesis. J. Mater. Chem. B 2025, 13, 858. https://doi.org/10.1039/D4TB01890G DOI: https://doi.org/10.1039/D4TB01890G
A. Precupas, I. Matei. Thioflavin T: induced circular dichroism and density functional theory as promising approach in amyloid fibril assays. Appl. Spectrosc. Rev., 2025, 61, 78. https://doi.org/10.1080/05704928.2025.2501692 DOI: https://doi.org/10.1080/05704928.2025.2501692
F. B. Cougnon, A. R. Stefankiewicz, S. Ulrich. Dynamic covalent synthesis. Chem. Sci., 2024, 15, 879. https://doi.org/10.1039/D3SC05343A DOI: https://doi.org/10.1039/D3SC05343A
Y. Chen, H. Tang, H. Chen, H. Li. Self-assembly via condensation of imine or its N-substituted derivatives. Acc. Chem. Res. 2023, 56, 2838. https://doi.org/10.1021/acs.accounts.3c00475 DOI: https://doi.org/10.1021/acs.accounts.3c00475
P. K. Kanchi, A. K. Dasmahapatra. Destabilization of the Alzheimer’s amyloid-β peptide by a proline-rich β-sheet breaker peptide: a molecular dynamics simulation study. J. Mol. Model., 2021, 27, 356 https://doi.org/10.1007/s00894-021-04968-x DOI: https://doi.org/10.1007/s00894-021-04968-x
N. Ghosh, L. M. Kundu. Breaker peptides against amyloid-β aggregation: a potential therapeutic strategy for Alzheimer’s disease. Future Med. Chem., 2021, 13, 1767. https://doi.org/10.4155/fmc-2021-0184 DOI: https://doi.org/10.4155/fmc-2021-0184
P. Ghosh, A. Bera, A. Ghosh, P. Bhadury, P. De. Side-chain proline-based polymers as effective inhibitors for in vitro aggregation of insulin. ACS Appl. Bio Mater., 2020, 3, 5407. https://doi.org/10.1021/acsabm.0c00709 DOI: https://doi.org/10.1021/acsabm.0c00709
A. Jarmuła, J. Ludwiczak, D. Stępkowski. β‐sheet breakers with consecutive phenylalanines: Insights into mechanism of dissolution of β‐amyloid fibrils. Proteins:Struct., Funct., Bioinf., 2021, 89, 762. https://doi.org/10.1002/prot.26057 DOI: https://doi.org/10.1002/prot.26057
K. Nayak, P. Ghosh, S. Barman, B. Sudhamalla, P. Theato, P. De. Amyloid β-Peptide Segment Conjugated Side-Chain Proline-Based Polymers as Potent Inhibitors in Lysozyme Amyloidosis. Bioconjug. Chem., 2024, 35, 312. https://doi.org/10.1021/acs.bioconjchem.3c00509 DOI: https://doi.org/10.1021/acs.bioconjchem.3c00509
I. J. Angera, M. M. Wright, J. R. Del Valle. Beyond N-Alkylation: Synthesis, Structure, and Function of N-Amino Peptides. Acc. Chem. Res., 2024, 57, 1287. https://doi.org/10.1021/acs.accounts.4c00024 DOI: https://doi.org/10.1021/acs.accounts.4c00024
M. Konar, D. Ghosh, S. Samanta, T. Govindaraju. Combating amyloid-induced cellular toxicity and stiffness by designer peptidomimetics. RSC Chem. Biol., 2022, 3, 220. https://doi.org/10.1039/D1CB00235J DOI: https://doi.org/10.1039/D1CB00235J
V. Apostolopoulos, J. Bojarska, T.-T. Chai, S. Elnagdy, K. Kaczmarek, J. Matsoukas, R. New, K. Parang, O. P. Lopez, H. Parhiz. A global review on short peptides: frontiers and perspectives. Molecules 2021, 26, 430. https://doi.org/10.3390/molecules26020430 DOI: https://doi.org/10.3390/molecules26020430
C. Lamers. Overcoming the shortcomings of peptide-based therapeutics. Future Drug Discov., 2022, 4, FDD75. https://doi.org/10.4155/fdd-2022-0005 DOI: https://doi.org/10.4155/fdd-2022-0005
S. M. P. Vadevoo, S. Gurung, H.-S. Lee, G. R. Gunassekaran, S.-M. Lee, J.-W. Yoon, Y.-K. Lee, B. Lee. Peptides as multifunctional players in cancer therapy. Exp Mol Med 2023, 55, 1099. https://doi.org/10.1038/s12276-023-01016-x DOI: https://doi.org/10.1038/s12276-023-01016-x
P. A. Valiente, H. Wen, S. Nim, J. Lee, H. J. Kim, J. Kim, A. Perez-Riba, Y. P. Paudel, I. Hwang, K.-D. Kim. Computational design of potent D-peptide inhibitors of SARS-CoV-2. J. Med. Chem., 2021, 64, 14955. https://doi.org/10.1021/acs.jmedchem.1c00655 DOI: https://doi.org/10.1021/acs.jmedchem.1c00655
K. S. Chen, K. Menezes, J. B. Rodgers, D. M. O’Hara, N. Tran, K. Fujisawa, S. Ishikura, S. Khodaei, H. Chau, A. Cranston. Small molecule inhibitors of α-synuclein oligomers identified by targeting early dopamine-mediated motor impairment in C. elegans. Mol. Neurodegener., 2021, 16, 77. https://doi.org/10.1186/s13024-021-00497-6 DOI: https://doi.org/10.1186/s13024-021-00497-6
S. Menon, S. Armstrong, A. Hamzeh, N. P. Visanji, S. P. Sardi, A. Tandon. Alpha-synuclein targeting therapeutics for Parkinson's disease and related synucleinopathies. Front. Neurol., 2022, 13, 852003. https://doi.org/10.3389/fneur.2022.852003 DOI: https://doi.org/10.3389/fneur.2022.852003
L. Ferrazzano, M. Catani, A. Cavazzini, G. Martelli, D. Corbisiero, P. Cantelmi, T. Fantoni, A. Mattellone, C. De Luca, S. Felletti. Sustainability in peptide chemistry: current synthesis and purification technologies and future challenges. Green Chem., 2022, 24, 975. https://doi.org/10.1039/D1GC04387K DOI: https://doi.org/10.1039/D1GC04387K
M. Zuo, T. Li, H. Feng, K. Wang, Y. Zhao, L. Wang, X. Y. Hu. Chaperone Mimetic Strategy for Achieving Organic Room‐Temperature Phosphorescence based on Confined Supramolecular Assembly. Small 2024, 20, 2306746. https://doi.org/10.1002/smll.202306746 DOI: https://doi.org/10.1002/smll.202306746
F. Siddiqui, P. Mishra, S. Khanam, S. Ranjan, P. Alam, T. Albalawi, S. Khan, S. S. Mir. Nano‐Chaperones: Bridging Therapeutics for Amyloid Aggregation in Alzheimer's Disease and Type‐2 Diabetes Mellitus. Eur. J. Neurosci., 2025, 61, e70142. https://doi.org/10.1111/ejn.70142 DOI: https://doi.org/10.1111/ejn.70142
A. Bera, S. Sahoo, K. Goswami, S. K. Das, P. Ghosh, P. De. Modulating Insulin Aggregation with Charge Variable Cholic Acid-Derived Polymers. Biomacromolecules 2021, 22, 4833. https://doi.org/10.1021/acs.biomac.1c01107 DOI: https://doi.org/10.1021/acs.biomac.1c01107
B. Joseph, S. Thomas, N. Sen, A. Paschold, W. H. Binder, S. Kumar. Bioinspired synthetic polymers-based inhibitors of Alzheimer's amyloid-β peptide aggregation. Polym. Chem., 2023, 14, 392. https://doi.org/10.1039/D2PY01217K DOI: https://doi.org/10.1039/D2PY01217K
P. Ghosh, A. Mondal, P. De. Polymeric Materials as a Promising Platform for Suppressing Protein Aggregation Process. Curr. Indian Sci., 2024, 2, 1. https://doi.org/10.2174/012210299X290674240509111832 DOI: https://doi.org/10.2174/012210299X290674240509111832
A. Mondal, M. E. H. Khan, P. Ghosh, P. De. Future Direction of Designing Antioxidant Polymers in Modulating Protein Aggregation Process. J. Mol. Eng. Mater., 2021, 09, 2140001. https://doi.org/10.1142/S2251237321400013 DOI: https://doi.org/10.1142/S2251237321400013
T. K. Paira, S. Banerjee, T. K. Mandal. Peptide-poly(ε-caprolactone) biohybrids by grafting-from ring-opening polymerization: Synthesis, aggregation, and crystalline properties. J. Polym. Sci. A Polym. Chem., 2012, 50, 2130. https://doi.org/10.1002/pola.26003 DOI: https://doi.org/10.1002/pola.26003
A. Saha, T. K. Paira, M. Biswas, S. Jana, S. Banerjee, T. K. Mandal. Combined atom-transfer radical polymerization and ring-opening polymerization to design polymer–polypeptide copolymer conjugates toward self-aggregated hybrid micro/nanospheres for dye encapsulation. J. Polym. Sci. A Polym. Chem., 2015, 53, 2313. https://doi.org/10.1002/pola.27713 DOI: https://doi.org/10.1002/pola.27713
T. K. Paira, A. Saha, S. Banerjee, T. Das, P. Das, N. R. Jana, T. K. Mandal. Fluorescent Amphiphilic PEG-Peptide-PEG Triblock Conjugate Micelles for Cell Imaging. Macromol. Biosci., 2014, 14, 929. https://doi.org/10.1002/mabi.201400083 DOI: https://doi.org/10.1002/mabi.201400083
E.-J. Nam, Y. Kwon, Y. Ha, S. R. Paik. Fabrication of a dual stimuli-responsive assorted film comprising magnetic-and gold-nanoparticles with a self-assembly protein of α-synuclein. ACS Appl. Bio Mater., 2021, 4, 1863. https://doi.org/10.1021/acsabm.0c01539 DOI: https://doi.org/10.1021/acsabm.0c01539
R. Boyuklieva, N. Zahariev, P. Simeonov, D. Penkov, P. Katsarov. Next-Generation Drug Delivery for Neurotherapeutics: The Promise of Stimuli-Triggered Nanocarriers. Biomedicines 2025, 13, 1464. https://doi.org/10.3390/biomedicines13061464 DOI: https://doi.org/10.3390/biomedicines13061464
X. j. Dai, W. J. Li, D. D. Xie, B. x. Liu, L. Gong, H. H. Han. Stimuli‐Responsive Nano Drug Delivery Systems for the Treatment of Neurological Diseases. Small 2025, 21, 2410030. https://doi.org/10.1002/smll.202410030 DOI: https://doi.org/10.1002/smll.202410030
A. Bera, P. Ghosh, S. Ghosh, A. Mukherjee, P. De. Antioxidant Polymers with Enhanced Neuroprotection Against Insulin Fibrillation. Macromol. Biosci., 2023, 23, 2300100. https://doi.org/10.1002/mabi.202300100 DOI: https://doi.org/10.1002/mabi.202300100
J. Basu, A. Soni, C. A. Athale. Physical effects of crowdant size and concentration on collective microtubule polymerization. Biophys. J., 2025, 124, 789. https://doi.org/10.1016/j.bpj.2025.01.020 DOI: https://doi.org/10.1016/j.bpj.2025.01.020
R. A. Norman, F. Ambrosetti, A. M. J. J. Bonvin, L. J. Colwell, S. Kelm, S. Kumar, K. Krawczyk. Computational approaches to therapeutic antibody design: established methods and emerging trends. Brief. Bioinform., 2019, 21, 1549. https://doi.org/10.1093/bib/bbz095 DOI: https://doi.org/10.1093/bib/bbz095